 
For the lagging strand, the direction of nucleotide polymerization is opposite to the overall direction of DNA chain growth. Its synthesis slightly precedes the synthesis of the daughter strand that is synthesized discontinuously, known as the lagging strand. The DNA daughter strand that is synthesized continuously is known as the leading strand. (Similar replication intermediates were later found in eucaryotes, where they are only 100–200 nucleotides long.) The Okazaki fragments were shown to be polymerized only in the 5′-to-3′chain direction and to be joined together after their synthesis to create long DNA chains.Ī replication fork therefore has an asymmetric structure ( Figure 5-8). 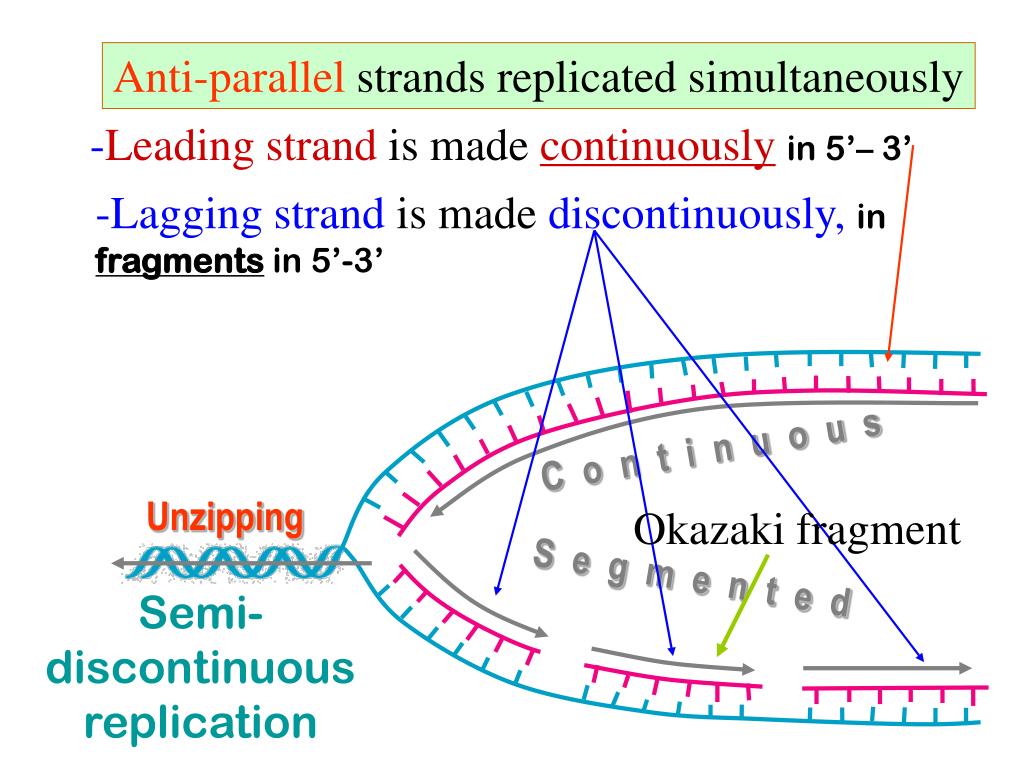 
This experiment revealed the transient existence of pieces of DNA that were 1000–2000 nucleotides long, now commonly known as Okazaki fragments, at the growing replication fork. Researchers added highly radioactive 3H-thymidine to dividing bacteria for a few seconds, so that only the most recently replicated DNA-that just behind the replication fork-became radiolabeled. How, then, is overall 3′-to-5′ DNA chain growth achieved? The answer was first suggested by the results of experiments in the late 1960s. In this scheme, both daughter DNA strands would grow continuously, using (more.) Although it might seem to be the simplest possible model for DNA replication, the mechanism illustrated here is not the one that cells use. 89–90), it does not occur in DNA synthesis no 3′-to-5′ DNA polymerase has ever been found.Īn incorrect model for DNA replication. Although head-growth polymerization occurs elsewhere in biochemistry (see pp. The other would move in the 3′-to-5′ direction and work by so-called “head growth,” in which the end of the growing DNA chain carried the triphosphate activation required for the addition of each subsequent nucleotide ( Figure 5-7). 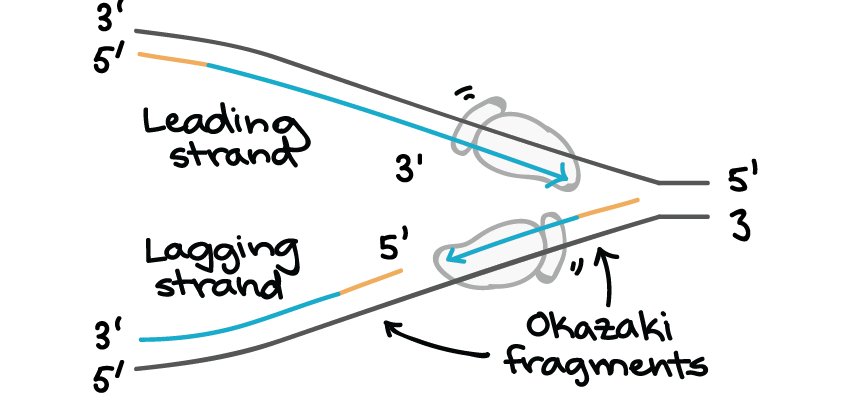
One would polymerize in the 5′-to-3′ direction, where each incoming deoxyribonucleoside triphosphate carried the triphosphate activation needed for its own addition. Such a replication fork would require two different DNA polymerase enzymes. But because of the antiparallel orientation of the two DNA strands in the DNA double helix (see Figure 5-2), this mechanism would require one daughter strand to polymerize in the 5′-to-3′ direction and the other in the 3′-to-5′ direction. Initially, the simplest mechanism of DNA replication seemed to be the continuous growth of both new strands, nucleotide by nucleotide, at the replication fork as it moves from one end of a DNA molecule to the other. An active zone of DNA replication moves progressively along a replicating DNA molecule, creating a Y-shaped DNA structure known as a replication fork: the two arms of each Y (more.) See also: DNA, replication, lagging strand.Two replication forks moving in opposite directions on a circular chromosome. Word origin: named after its discoverers, Reiji Okazaki and his wife, Tsuneko Okazaki, while studying replication of bacteriophage DNA in Escherichia coli in 1968. Okazaki fragments are originally discovered by Reiji Okazaki, Tsuneko Okazaki, and their colleagues while studying replication of bacteriophage DNA in Escherichia coli in 1968. This is because DNA synthesis can proceed only in one direction - the 5′ to 3′ direction. Unlike the leading strand where DNA can be synthesized continuously the lagging strand is synthesized discontinuously in the form of short fragments called Okazaki fragments that are later connected covalently to form a continuous strand. One of the strands goes from 5’ to 3’ and is called the leading strand the other strand goes from a 3′ to 5′ and is called the lagging strand. Relatively short fragment of DNA synthesized on the lagging strand during DNA replication.Īt the start of DNA replication, DNA unwinds and the two strands splits in two, forming two “prongs” which resemble a fork (thus, called replication fork). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
